Researchers from CUNY ISPH and colleagues recently published a powerful new data structure for the analysis of genomic data for open-source statistical computing.
In genomic research, scientists analyze various aspects of DNA, such as copy number, mutation and chemical modifications, to understand how genes function and contribute to diseases like cancer. However, the data generated from these experiments present informatics challenges to overcome before any statistical analyses can be performed: like a puzzle whose pieces don’t fit neatly together, each sample has observations at different genomic locations.
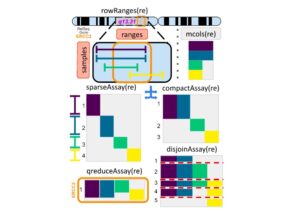
To address this challenge, CUNY SPH alum and Senior Data Scientist Marcel Ramos, ISPH Investigator Levi Waldron and colleagues from the Harvard T.H. Chan School of Public Health, Harvard Medical School and the Roswell Park Comprehensive Cancer Center developed a new approach called RaggedExperiment in the R/Bioconductor statistical programming environment. It allows for organized representation of this “ragged” genomic data, preserving all the information and providing tools that make it easier to transform and analyze such data in different ways.
“There has been no Bioconductor data class for lossless representation of ragged genomic data within the Bioconductor ecosystem of packages for multi-omic data analysis, or to facilitate flexible conversion to matrix representations such as number of coding mutations or copy number per gene,” says Ramos. “RaggedExperiment adds a more powerful, efficient, and less error-prone tool to the genomic data analyst’s toolbox.”
“Marcel has developed and refined this software over several years and it has already found a significant user base, so I’m really pleased to formally describe and publish it in one of the top journals in the field of Bioinformatics,” says Waldron.“By enhancing our ability to analyze and understand genomic data, this development opens up new possibilities for improving our knowledge of diseases and developing better treatments.”
The RaggedExperiment package is publicly available under an Artistic 2.0 license from the Bioconductor project for open-source Bioinformatics, with open development and issue tracking on GitHub.
Marcel Ramos, Martin Morgan, Ludwig Geistlinger, Vincent J. Carey, Levi Waldron. RaggedExperiment: the missing link between genomic ranges and matrices in Bioconductor. Bioinformatics. 2023 May 19:btad330. doi: 10.1093/bioinformatics/btad330. Epub ahead of print. PMID: 37208161.